aslprep.workflows.asl.confounds module
Workflows for calculating confounds for ASL data.
- init_asl_confounds_wf(n_volumes: int, mem_gb: float, freesurfer: bool = False, name: str = 'asl_confounds_wf')[source]
Build a workflow to generate and write out confounding signals.
This workflow calculates confounds for a asl series, and aggregates them into a TSV file, for use as nuisance regressors in a GLM. The following confounds are calculated, with column headings in parentheses:
DVARS - original and standardized variants (
dvars,std_dvars)Framewise displacement, based on head-motion parameters (
framewise_displacement)Estimated head-motion parameters, in mm and rad (
trans_x,trans_y,trans_z,rot_x,rot_y,rot_z)
- Workflow Graph
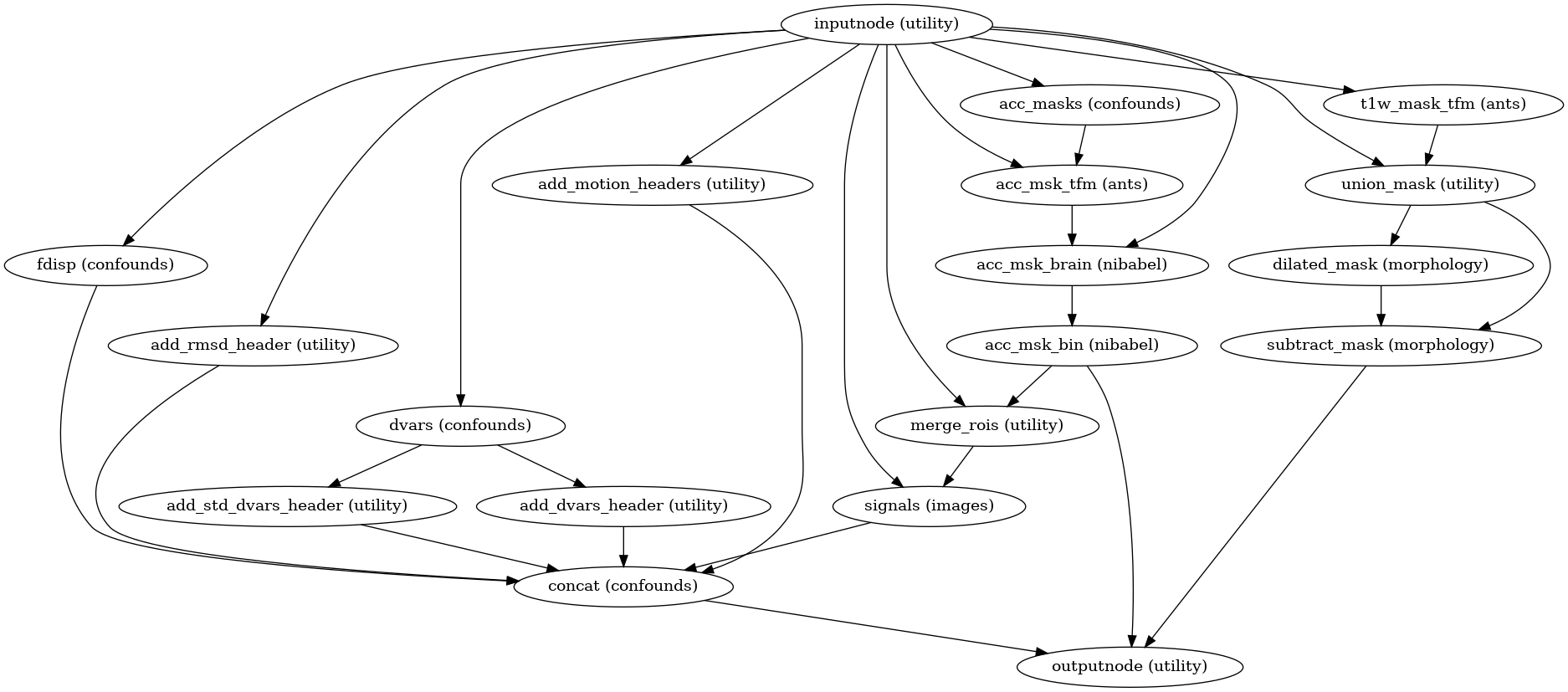
(Source code, png, svg, pdf)
- Parameters:
n_volumes (
int) – Number of volumes in the ASL file. Some relative measures (e.g., FD, DVARS, RMSD) will produce empty arrays for single-volume datasets, so we must skip those.mem_gb (
float) – Size of asl file in GB - please note that this size should be calculated after resamplings that may extend the FoVfreesurfer (
bool) – Set toTrueif the input volume fractions for the anatomical component-based noise correction maps come from FreeSurfer’saseg. sMRIPrep always uses FAST for tissue probability maps, so we always set this to False.name (
str) – Name of workflow (default:asl_confounds_wf)
- Inputs:
asl – asl image, after the prescribed corrections (HMC and SDC) when available.
asl_mask – asl series mask
hmc_aslref – ASL reference image used for HMC
coreg_aslref – ASL reference image used for coregistration
motion_xfm – ITK-formatted head motion transforms
t1w_mask – Mask of the skull-stripped template image
t1w_tpms – List of tissue probability maps in T1w space
aslref2anat_xfm – Affine matrix that maps the T1w space into alignment with the native asl space
- Outputs:
confounds_file – TSV of all aggregated confounds
confounds_metadata – Confounds metadata dictionary.
crown_mask – Mask of brain edge voxels
acompcor_masks
- init_carpetplot_wf(mem_gb: float, confounds_list: list, metadata: dict, cifti_output: bool, suffix: str = 'asl', name: str = 'asl_carpet_wf')[source]
Build a workflow to generate carpet plots.
Resamples the MNI parcellation (ad-hoc parcellation derived from the Harvard-Oxford template and others).
- Parameters:
- Inputs:
asl – ASL image, after the prescribed corrections (STC, HMC and SDC) when available.
asl_mask – ASL series mask
confounds_file – TSV of all aggregated confounds
aslref2anat_xfm – Affine matrix that maps the T1w space into alignment with the native ASL space
std2anat_xfm – ANTs-compatible affine-and-warp transform file
cifti_asl – ASL image in CIFTI format, to be used in place of volumetric ASL
crown_mask – Mask of brain edge voxels
acompcor_mask – Mask of deep WM+CSF
- Outputs:
out_carpetplot – Path of the generated SVG file
- init_cbf_confounds_wf(scorescrub=False, basil=False, name='cbf_confounds_wf')[source]
Create a workflow for dolui2024automated.
- Workflow Graph
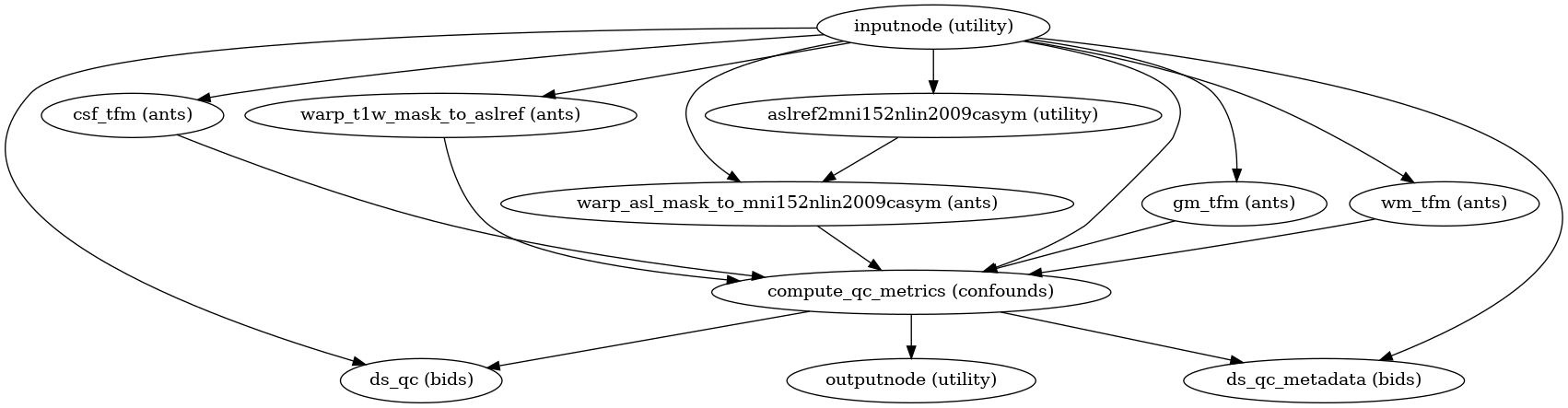
(Source code, png, svg, pdf)
- Parameters:
scorescrub (bool)
basil (bool)
name (
str) – Name of workflow (default: “cbf_qc_wf”)
- Inputs:
*cbf – all cbf
asl_mask – asl mask NIFTI file
t1w_tpms – t1w probability maps
aslref2anat_xfm – aslref to t1w transformation file
anat2mni2009c_xfm
asl_mask_std (list) – Since ASLPrep always includes MNI152NLin2009cAsym as a standard space, this should always be provided.
- Outputs:
qc_file – qc measures in tsv